CRAN release ‘Luminescence’ v0.9.19
- the lazy cat wants peace -
by RLum.Network (March 11, 2022)
Finally! 'Luminescence' was the last package that needed an update
after the CRAN request
had triggered an unscheduled number of releases in a short period.
Thanks to the always-active CRAN team, 0.9.19 made it on CRAN
yesterday night.
Technique babel
Then the last check prior to submission revealed some additional problems that may
probably haunt us for some time in the future. For performance reasons, we had used if(class(object) == "something"))
multiple times in nearly all functions, instead of other solutions:
microbenchmark::microbenchmark(
is = is("test", "character"),
inhertis = inherits("test", "character"),
class_sub = class("test")[1] == "character",
class = class("test") == "character")## Unit: nanoseconds
## expr min lq mean median uq max neval cld
## is 6611 6751.0 7101.54 6842.5 6984.0 26364 100 c
## inhertis 650 690.0 767.04 732.0 777.5 3189 100 b
## class_sub 373 405.5 474.22 459.0 502.5 1610 100 ab
## class 234 254.5 319.43 269.0 300.0 4537 100 aThe newest R-devel checks now flagged this construction as problematic.
Well, admittedly, class(object) was ill-chosen in the first place because it was not justified to expect the output to be a vector of length 1.
This means the solution was error-prone and sanitised by class(object)[1]. Unfortunately,
a couple of wrong code (too many) remained and needed a replacement. Given the poor performance
of is(), we opted for inherits().
This solution may probably lead to a marginal slower performance while sub-setting large objects; OK.
New functions
Besides these non-user-visible changes under the hood, the new release brings two
new functions, adding to the skill set of 'Luminescence'.
extract_ROI()
This function allows you to extract circular regions-of-interest from image data.
While other software solutions do image processing way fast and better, in the context
of luminescence data analysis, this something for which you don’t want to swap
software. While in the example below, the function uses a self-created matrix as an image
input, array and RLum.Data.Image objects are other possibilities.
m <- matrix(runif(100,0,255), ncol = 10, nrow = 10)
roi <- matrix(c(2.,4,2,5,6,7,3,1,1), ncol = 3)
extract_ROI(object = m, roi = roi, plot = TRUE)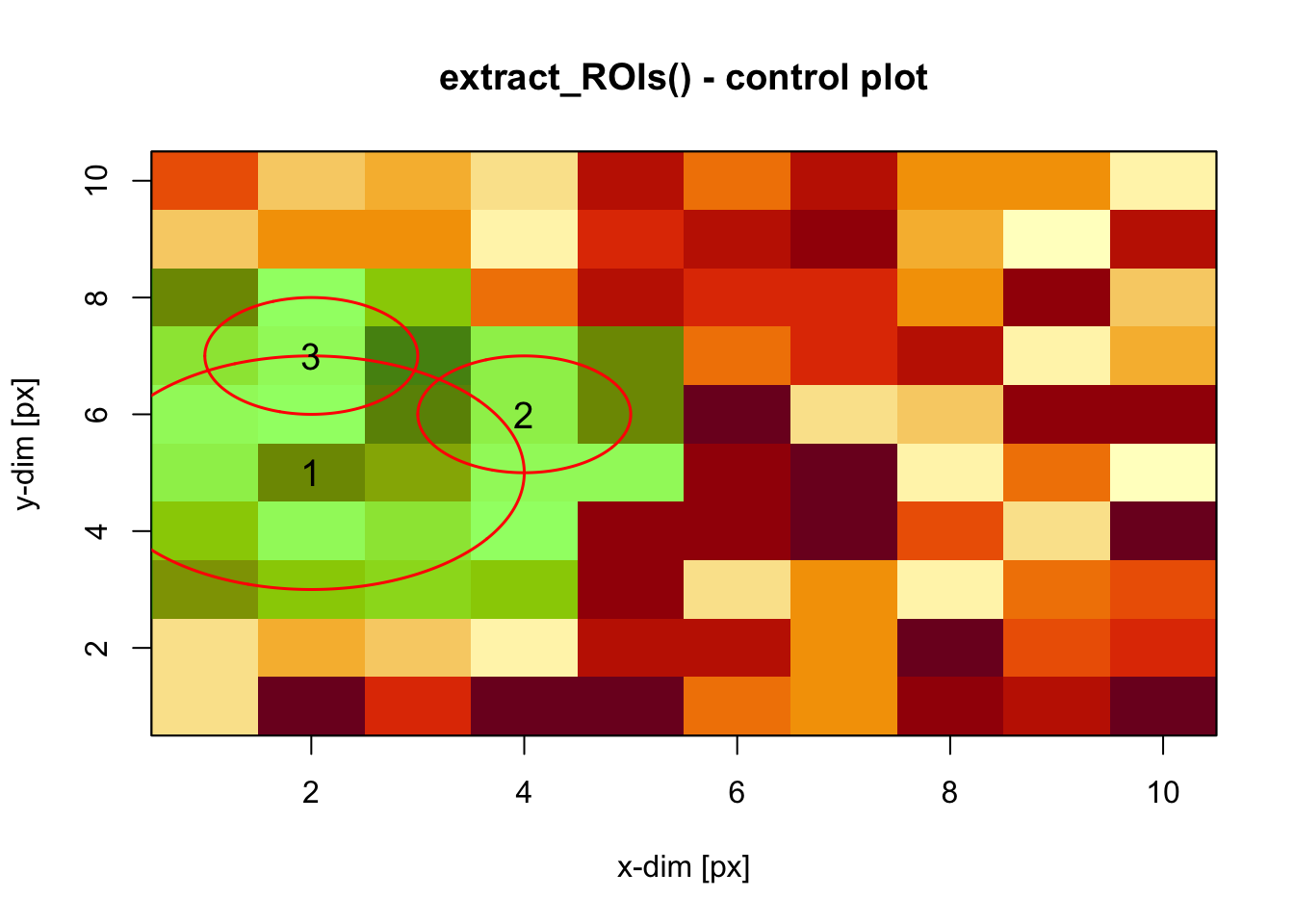
##
## [RLum.Results-class]
## originator: extract_ROI()
## data: 3
## .. $roi_signals : list
## .. $roi_summary : matrix
## .. $roi_coord : matrix
## additional info elements: 1write_R2TIFF()
It was always somewhat annoying that image and spectrum data could not be
processed in R and then returned to an image file. write_R2TIFF() solves
that problem.
Further news?
Small fixes, minor changes, nothing out of the ordinary. For a complete list of changes, please check our GitHub page.
Stay safe and enjoy 'Luminescence'
Sebastian on behalf of the R Luminescence Team